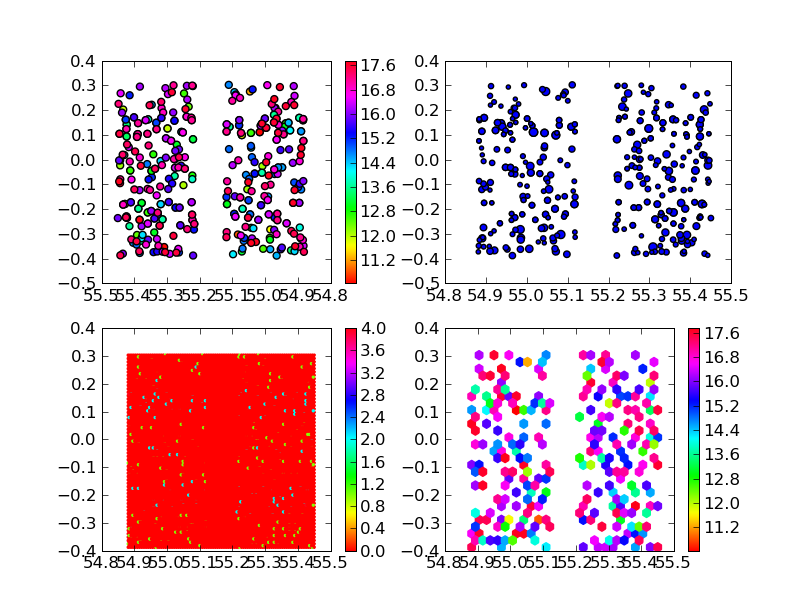
Example data for making these plots. If you want to reproduce these plots with the commands above, first do
The lower two subplots show the easiest way to do spatial binning, with hexbin. The lower left plot shows just the number of data points in each hexagonal cell (zero for most cells, hence the monotonous red), using
hexbin(ra,dec,cmap=cm.hsv) colorbar()The lower right plot shows how to average some other quantity in the hexagonal cells, with
hexbin(ra,dec,C=mag,gridsize=25,cmap=cm.hsv) colorbar()The number of hex cells is now reduced from the default 100, so that I sometimes get more than one data point in a cell! The absence of data is shown at white; I probably should have used a denser data set for this example. If you don't like hexagonal bins, scroll down to "More Spatial Binning" to see an alternative, which takes more than one line of code.
To do outlier rejection, try plotting the median rather than the mean of the quantities in each cell: hexbin(ra,dec,C=mag,gridsize=25,cmap=cm.hsv,reduce_C_function=numpy.median))

Example data for making these plots. If
you want to reproduce these plots with the commands above, first do
data = numpy.loadtxt('example.fiat')
ra = data[:,3]
dec = data[:,4]
mag = data[:,5]
(after your normal import statements, of course). Note: on very old
installations of matplotlib, the hexbin function does not exist. If
you need it, update your system!
The way I prefer to accommodate crowded scatterplots, which I inherited from sm, the plotting package I used prior to matplotlib, is to make points "open," that is, they have a perimeter but nothing in the center so that it is easy to see exactly how markers overlap. In matplotlib this is accomplished with
plot(v1, v2, 'o', markerfacecolor='None')Before a reader told me about this solution, my workaround was to keep the markers big enough to overlap, but make them partially transparent with the option alpha=0.2 or so.
In summary, you can separately control the color of the face and the perimeter of the marker, including by specifying None for either color. This was not obvious to me just by looking at the example gallery.
A challenge for matplotlib experts out there: how do I make a plot which is a scatterplot when the density of points is low, but transitions to a contour plot when the density of points is high? I've seen these types of plot prepared by other graphics systems, and they seem to me to be the optimal way to represent density when both high and low density regions are interesting.
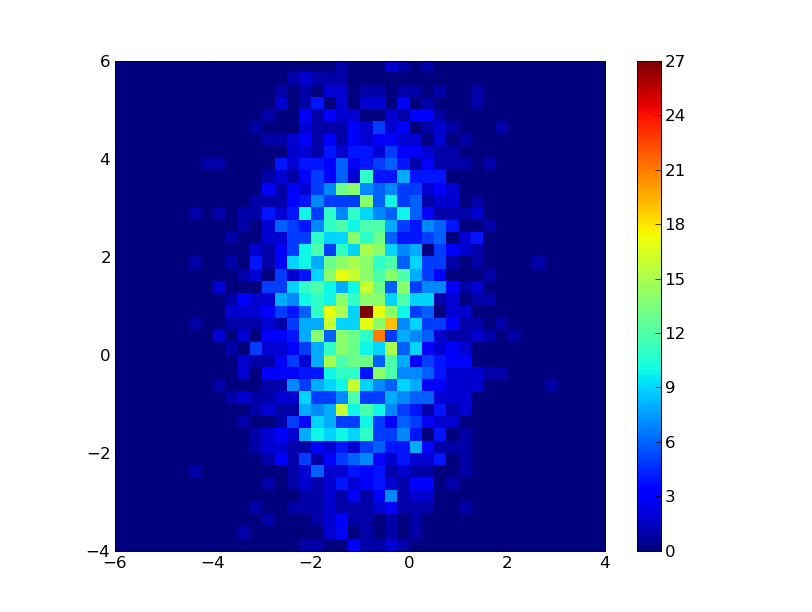
Here is the simplest test case. It generates Gaussian distributions with different offsets and variances in x and y so you can verify that it's really doing the right thing. Notice the use of the transpose, hist.T.
from pylab import * import numpy # the x distribution will be centered at -1, the y distro # at +1 with twice the width. x = numpy.random.randn(3000)-1 y = numpy.random.randn(3000)*2+1 hist,xedges,yedges = numpy.histogram2d(x,y,bins=40,range=[[-6,4],[-4,6]]) extent = [xedges[0], xedges[-1], yedges[0], yedges[-1] ] imshow(hist.T,extent=extent,interpolation='nearest',origin='lower') colorbar() show()On my machine at least, it produces this image:
maps=[m for m in cm.datad if not m.endswith("_r")]
print maps
Each (some?) map name also has a _r version to reverse it.
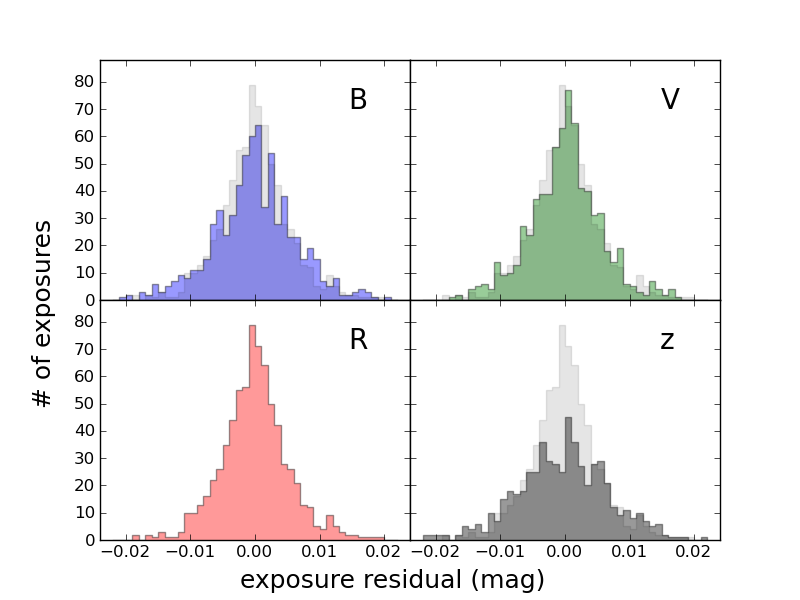
from matplotlib.ticker import NullFormatter ... ax = subplot(222) ax.xaxis.set_major_formatter( NullFormatter() ) ax.yaxis.set_major_formatter( NullFormatter() )You may also need to adjust the locations of the labels. In the above example, matplotlib wanted to put labels at every 0.025 on the x axes, so the +0.025 in one panel collided with the -0.025 in the adjacent panel. So I used
ax.xaxis.set_major_locator(MultipleLocator(0.01))To make it clear that the label "exposure residual (mag)" applies to all subplots, I used figtext which places text in overall figure coordinates (0-1 in each direction, from lower left) rather than xlabel, which places it relative to your current subplot.
Note that this plot also makes use of the alpha transparency option discussed briefly above. You can use alpha with lots of routines.
To see how all these bits fit together, read the full script used to make the above plot.
xlabel(string,fontdict={'fontsize':20})
Here is a comparison of a shrunken figure with and without this
optional argument: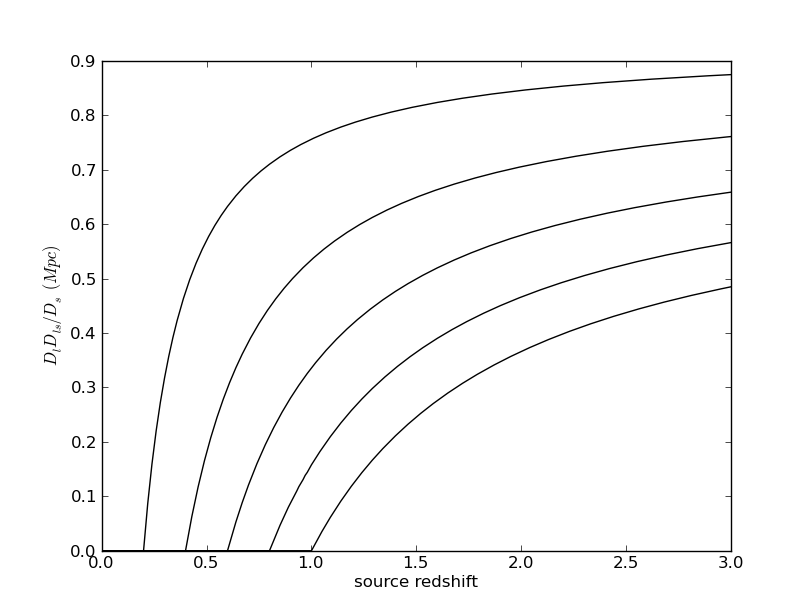
Fiddling with which ticks are labeled: sometimes the default choice of which ticks are labeled is awkward, for example when labels at the corners run into each other. You can control this with:
from matplotlib.ticker import MultipleLocator ... majorLocator = MultipleLocator(0.01) ax=subplot(223) ax.xaxis.set_major_locator( majorLocator )This puts a label every 0.01 units as in the exposure residual example above, in the Subplots section. In that plot, for consistency I used the same tick locator in multiple subplots.
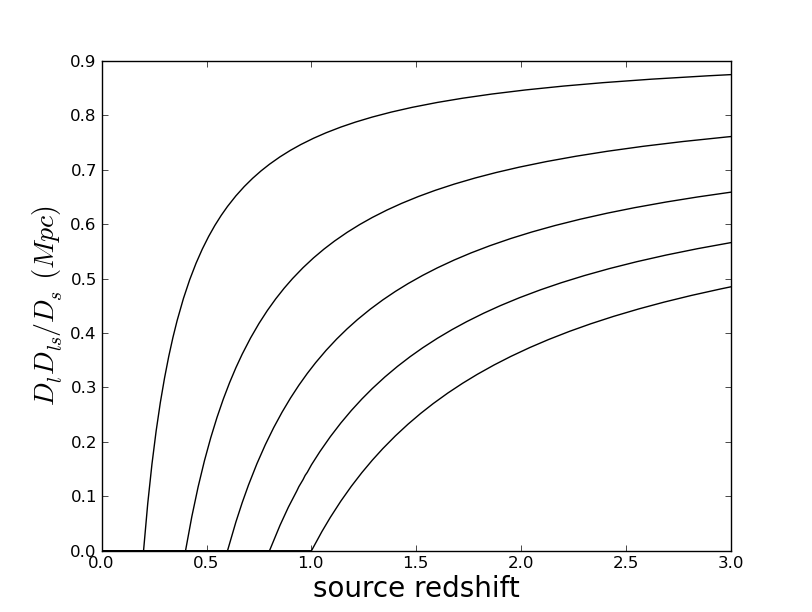
Math symbols in labels: everyone knows that you can put TeX in the labels, but it was not obvious to me how to quote a string to make this happen correctly. Here's how:
ylabel(r'$D_l D_{ls}/D_s\ (Mpc)$',fontdict={'fontsize':20})
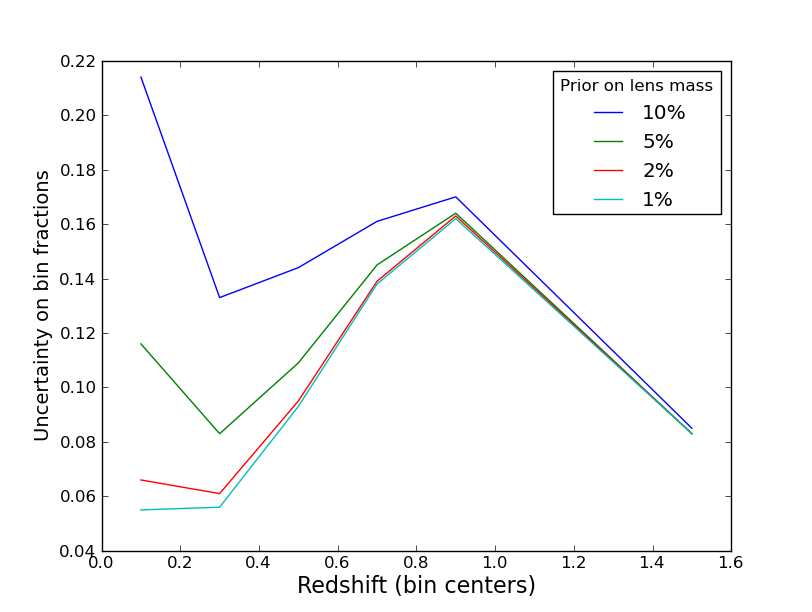
This is exactly the code used in the above plot.Legends: these are nice when plotting multiple curves, and are set up to work basically automatically. Here is the result of
for i in range(len(priors)):
plot(z,y[i,:],label='%d%%' % (priors[i]*100))
legend(title='Prior on lens mass')
plot((x1,x2),(y1,y2),'k-')This still doesn't work as nicely as sm's draw command, because it makes matplotlib think these are datapoints, and thus expands the plot if the points you give are outside the range of the other (real) data. So you have to pay more attention to the points you give. Being lazy, I would like a command that says "draw a line from this point to that point, but still use the range of real data to determine the cropping."